 Unfortunately, information in REMARK 350 is often incorrect (see discussion of this problem by Roland Dunbrack). This is given in REMARK 350 in the header of the PDB file format. When a structure is deposited in the PDB, the authors are required to specify the biological unit if it is known. Unreliability of REMARK 350 in the PDB File Header How To Show The Biological Unit In Proteopedia When there is more than one biological unit, all will be listed and you can display and analyze each of them. In the Molecule Information tab (the first/left-most tab), click Biological Unit and follow instructions.Alternatively, go directly to (google firstglance, one word with no space) and enter the PDB code.At the page in Proteopedia titled with the PDB code, under Resources, click on the link to FirstGlance. 
Enter the PDB code in the top search slot at the left edge of any page in Proteopedia.Display the molecule in FirstGlance in Jmol:.When it is very large, it will be simplified to alpha carbon atoms (or a subset thereof) automatically by FirstGlance. The initial display in FirstGlance is automatically "biomolecule 1" from REMARK 350. An example is the much larger capsid of Eastern Equine Encephalitis Virus, 6mx4.įirstGlance in Jmol makes it quick and easy to see, explore, and analyze the biological unit. When biological unit 1 becomes too large for JSmol to handle effectively, it is displayed in Mol* (" Molstar") instead of JSmol. A large example is 1pov, the polio virus capsid. A simple example is 3hyd: biological unit 1 has 4 small peptide chains, while the asymmetric unit has a single chain. On pages titled with a PDB code, Proteopedia shows biological unit 1 by default, with an option to show the asymmetric unit instead. 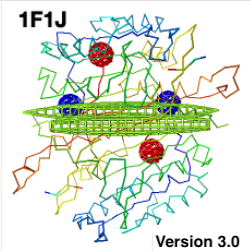
Visualizing the Biological Unit Proteopedia Mutation Y397D prevents this artifactual dimerization, leading to the monomer 1ee5. For example, 1bk5 (karyopherin alpha) is a truncated part of the natural chain, and forms a dimer that would be prevented by the full-length chain. Truncated proteins may form oligomers that are impossible in the native protein. **The "author specified" assembly (in this case the same as the asymmetric unit) appears unlikely in view of the assembly predicted by PISA, which has a much larger buried surface area. * The contacts in this biological unit differ from those in the asymmetric unit. In the PDB file format, this is done in REMARK 350. When publishing a macromolecular structure, the authors may elect to specify the biological unit. Interchain contacts that occur in the asymmetric unit that are absent in the biological unit are termed crystal contacts. Published macromolecular structure data files ( Atomic coordinate files, often in the PDB file format) contain the Asymmetric Unit, which may be identical with the biological unit, or only a portion of it, or may contain multiple biological units. For example, phosphorylation or dephosphorylation by protein kinases or phosphatases often change the affinities between proteins, and hence their quaternary assemblies. Of course, the functional form (biological unit) under one set of conditions may change under a different set of conditions, so there may be more than one functional form (biological unit) that includes a given protein chain. When a biological unit contains multiple chains that have co-evolved to bind to each other, it may also be referred to as a specific oligomer. For example, the biological unit of hemoglobin includes two alpha chains and two beta chains, making it a tetrameric α 2β 2 structure. It can be a single chain, or a quaternary assembly of multiple identical or non-identical chains. The Biological Unit, also called the Biological Assembly, is the quaternary structure of a protein that is believed to be the main functional form of the molecule.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
